|
To include the annotation, you need the sequence in Genbank format:Ĭopy the text and paste it in Notepad. On the next page select the Sequence tab: On the next page containing the full sequence of the vector, click the Analyze Sequence button on the right: On the webpage of mRFP-Rab5 go to the SEQUENCE INFORMATION section and click Full (1) next to Depositor Sequences: Obtain the annotated sequence of mRFP-Rab5 from Addgenes. If you load the annotations (in Genbank format) SnapGene will automatically show them. Genbank format not only contains a sequence but also all annotations. Even if you don't load them from Genbank it is still a good idea to import them in Genbank format (and not in Fasta format). In SnapGene you can load sequences directly from Genbank or you can load them by importing a file. Fortunately, many vectors are available from Addgene, an organization that improves sharing of plasmids. However, not all vector sequences are stored in the SnapGene database. Loading vector sequences from other resources Go back to SnapGene and click the Open button in the top menu:īrowse to the pAcGFP1-C1.dna file and open it. On the webpage of pAcGFP1-C1 click pAcGFP1-C1.dna:  
Try to load it without peeking at the solution. This vector is available in SnapGene's list of fluorescent protein genes and plasmids. You now see the circular view of the vector with restriction sites automatically visualized:Įxercise: load the sequence of vector pAcGFP1-C1 Open SnapGene and select Open in the startup menu:īrowse to the pcDNA4 TO.dna file and open it. Open the annotated sequence of pcDNA4/TO in SnapGene. On the webpage of pcDNA4/TO click the pcDNA4 TO.dna link to save the file: The application does not feature the functionality included in tools of the same feather, but it offers support for the most frequently used operations as far as DNA sequence analysis is concerned.Annotation: every possible piece of information about a sequenceĭownload the annotated sequence of pcDNA4/TO from SnapGene. Primers for PCR, sequencing, or mutagenesis can be designed and annotated, too. It sports automatic annotation of common features but it also offers the opportunity to do it manually in the case of coding sequences as well as more particular features. Features at a glanceĪmong the list of features available in SnapGene Viewer there is the possibility to create a DNA sequence file from punching in the sequence and export it to a GenBank format. There are multiple views available that allow toggling the display of enzymes on or off as well as showing the sequences, features or primers. However, a single look at the interface shows that the developers did their best to come up with a layout that is intuitive and comprehensive at the same time.Īs soon as a DNA file is loaded, there is a clear view of the map. User friendly interfaceĭue to the domain it has been built for, SnapGene is not accessible to all users. 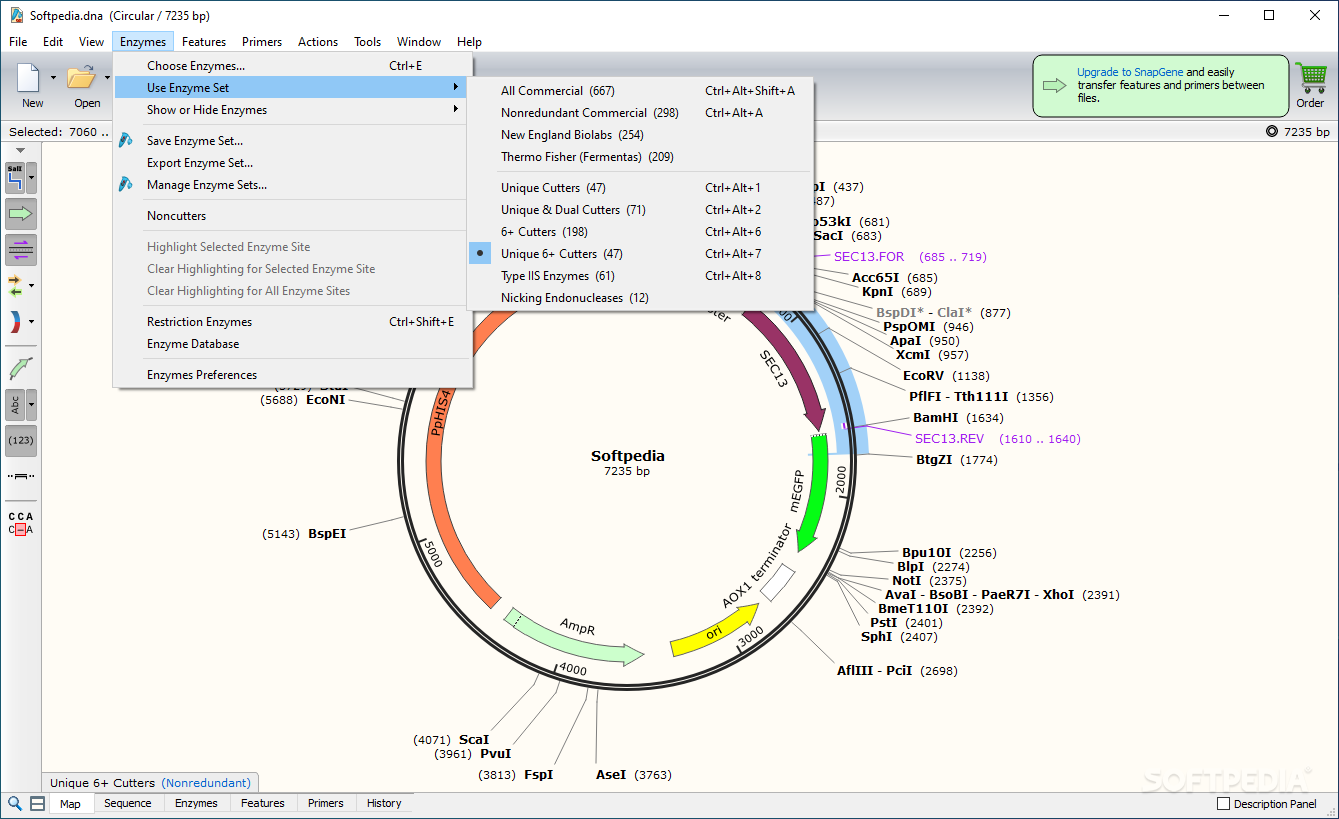
Getting the application on the system is done through an uneventful installation process that includes the option to associate specific file extensions (sequences, sequence traces and archives) with SnapGene Viewer. The application works with files as large as 1GB. SnapGene Viewer has been designed as a helpful tool for biologists to handle and exchange annotated DNA sequences easier and with less effort. Luckily, some developers provide the necessary digital instruments for analyzing this type of data in an easier way. Working with DNA sequences can be a difficult task, even for those who are familiar with the matter.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
